# total number of unique positions, indicating that several sites have two or more alternate allelesīcftools view -v snps NA12878.19.vcf.gz | grep -v "^#" | cut -f2 | sort -u | wc -l (I chose this file because it is relatively small: 4.1M.)īcftools view -v snps NA12878.19.vcf.gz | grep -v "^#" | wc -l I used BCFtools with some other command line tools to get a feel of the VCF file. I will use NA12878.19.vcf.gz as an example VCF file this file can be downloaded from the GATK bundle but I have also provided it in my GitHub repository. Using conda env create will build a Conda environment with all the required tools for this post. Below are the contents of variant.yml, which is also in my GitHub repository.
#BCFTOOLS MANUAL INSTALL#
# run the following commands inside DockerĬonda install -y -c bioconda perl-vcftools-vcfĪlternatively, you can prepare a YAML file that contains the required tools and create a Conda environment. Next, pull the Conda image and use Conda to install the various tools that will be used in this post.ĭocker run -rm -v /Users/dtang/github/learning_vcf_file:/data -i -t continuumio/miniconda /bin/bash If you don’t have Docker, install it first.
The repo also has a bunch of notes on VCF files in general. The example VCF file and other scripts used for this post are available in my GitHub repository, so please clone that somewhere if you want to follow this post. I have written some notes on Docker and Conda that maybe useful. I highly recommend learning about these tools if you haven’t already they make it easier to reproduce your work. To create a reproducible example, I will make use of Docker and Conda.

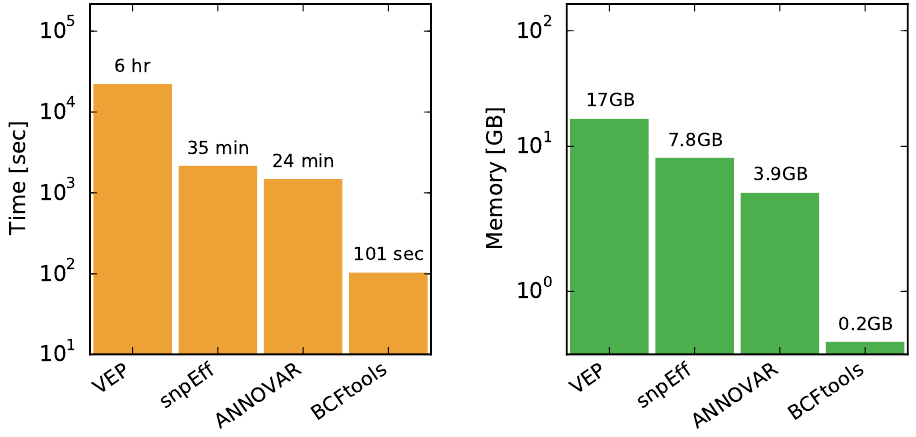
In this post, I will compare different tools for comparing VCF files.

